Note
Go to the end to download the full example code
PCA analysis example¶
In this example, we perform the PCA dimensionality reduction of the classical iris dataset (Ronald A. Fisher.
“The Use of Multiple Measurements in Taxonomic Problems. Annals of Eugenics, 7, pp.179-188, 1936).
First we laod the spectrochempy API package
import spectrochempy as scp
Upload a dataset form a distant server
try:
dataset = scp.download_iris()
except (IOError, OSError):
print("Could not load The `IRIS` dataset. Finishing here.")
import sys
sys.exit(0)
Create a PCA object
Reduce the data to a lower dimensionality. Here, the number of
components is automatically determined using n_pc="auto". As
indicated by the dimension of LT, 4 PC are found.
S, LT = pca.reduce(n_pc="auto")
print(LT)
NDDataset: [float64] cm (shape: (y:3, x:4))
The figures of merit (explained and cumulative variance) confirm that these 4 PC’s explain 100% of the variance:
PC Eigenvalue %variance %cumulative
of cov(X) per PC variance
#1 1.449e+01 92.462 92.462
#2 3.469e+00 5.302 97.763
#3 1.975e+00 1.719 99.482
#4 1.085e+00 0.518 100.000
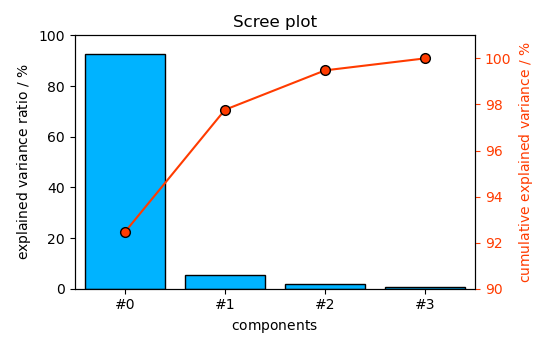
These figures of merit can also be displayed graphically
The ScreePlot
_ = pca.screeplot()
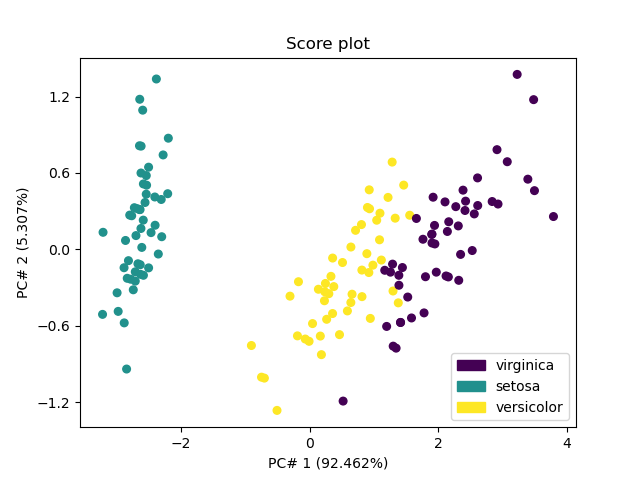
The score plots can be used for classification purposes. The first one - in 2D for the 2 first PC’s - shows that the first PC allows distinguishing Iris-setosa (score of PC#1 < -1) from other species (score of PC#1 > -1), while more PC’s are required to distinguish versicolor from viginica.
_ = pca.scoreplot(1, 2, color_mapping="labels")
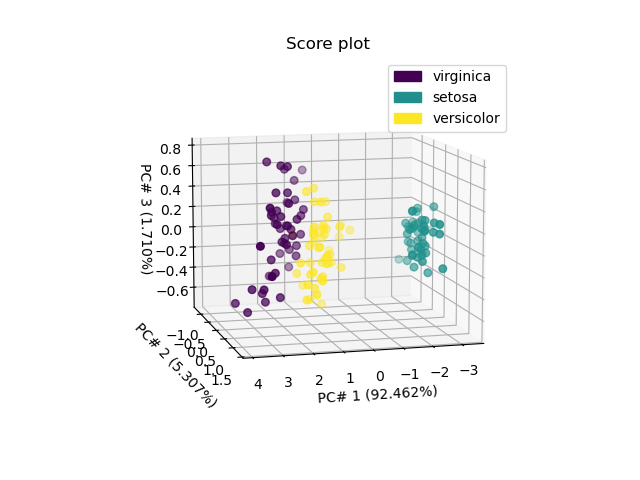
The second one - in 3D for the 3 first PC’s - indicates that a thid PC won’t allow better distinguishing versicolor from viginica.
ax = pca.scoreplot(1, 2, 3, color_mapping="labels")
ax.view_init(10, 75)
# scp.show() # uncomment to show plot if needed (not necessary in jupyter notebook)
Total running time of the script: ( 0 minutes 1.131 seconds)