Note
Go to the end to download the full example code
SIMPLISMA example¶
In this example, we perform the PCA dimensionality reduction of a spectra dataset
Import the spectrochempy API package (and the SIMPLISMA model independently)
import spectrochempy as scp
Load a matlab datasets
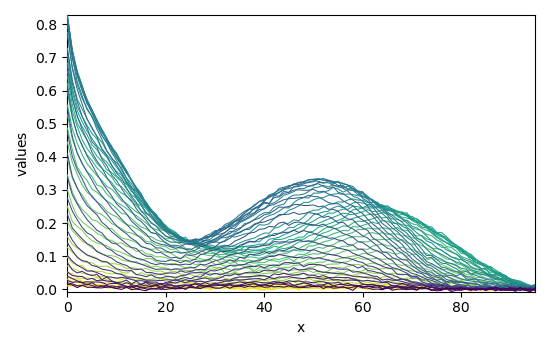
Dataset (Jaumot et al., Chemometr. Intell. Lab. 76 (2005) 101-110)):
cpure (204, 4)
MATRIX (204, 96)
isp_matrix (4, 4)
spure (4, 96)
csel_matrix (51, 4)
m1 (51, 96)
Add some metadata for a nicer display
ds.title = "absorbance"
ds.units = "absorbance"
ds.set_coordset(None, None)
ds.y.title = "elution time"
ds.x.title = "wavelength"
ds.y.units = "hours"
ds.x.units = "nm"
Fit the SIMPLISMA model
print("Fit SIMPLISMA on {}\n".format(ds.name))
simpl = scp.SIMPLISMA(max_components=20, tol=0.2, noise=3, log_level="INFO")
simpl.fit(ds)
Fit SIMPLISMA on m1
/home/runner/micromamba-root/envs/scpy/lib/python3.9/site-packages/spectrochempy/utils/decorators.py:85: DeprecationWarning: The `max_components` attribute is now deprecated. Use `n_components` instead. `max_components` attribute will be removed in version 0.7.
warn(
/home/runner/micromamba-root/envs/scpy/lib/python3.9/site-packages/spectrochempy/analysis/simplisma.py:190: UserWarning: SIMPLISMA does not handle easily negative values.
warn("SIMPLISMA does not handle easily negative values.")
<spectrochempy.analysis.simplisma.SIMPLISMA object at 0x7f15af9543d0>
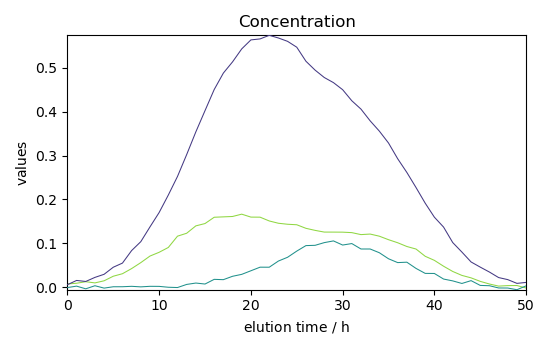
Plot concentration
_ = simpl.C.T.plot(title="Concentration")
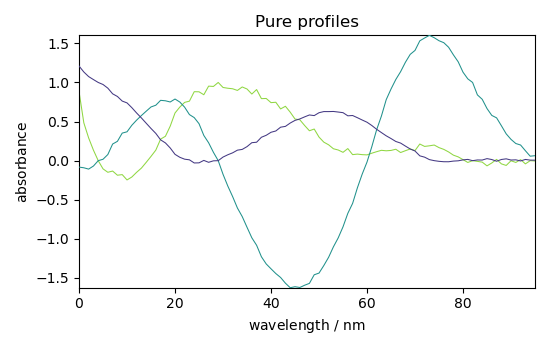
Plot components (St)
# sphinx_gallery_thumbnail_number = 3
_ = simpl.components.plot(title="Pure profiles")
Show the plot of merit after reconstruction oto the original data space
simpl.plotmerit(offset=0, nb_traces=5)
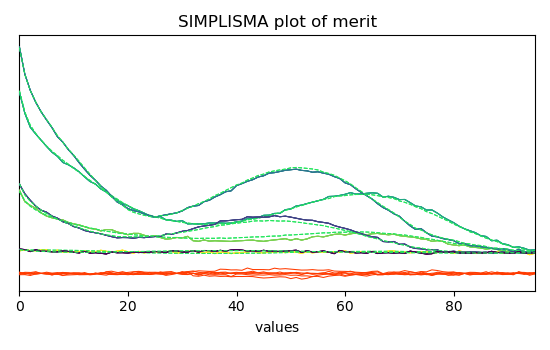
<_Axes: title={'center': 'SIMPLISMA plot of merit'}, xlabel='values $\\mathrm{}$', ylabel='absorbance $\\mathrm{/\\ \\mathrm{a.u.}}$'>
This ends the example ! The following line can be uncommented if no plot shows when running the .py script
# scp.show()
Total running time of the script: ( 0 minutes 0.887 seconds)