Note
Go to the end to download the full example code
NDDataset coordinates example¶
In this example, we show how coordinates can be used in SpectroChemPy
import spectrochempy as scp
Uploading a dataset¶
X has two coordinates:
* wavenumbers named “x”
* and timestamps (i.e., the time of recording) named “y”.
print(X.coordset)
CoordSet: [x:wavenumbers, y:acquisition timestamp (GMT)]
To display them individually use the x and y attributes of
the dataset X:
Setting new coordinates¶
In this example, each experiment have a timestamp corresponds to the time when a given pressure of CO in the infrared cell was set.
Hence, it would be interesting to replace the “useless” timestamps (y )
by a pressure coordinates:
pressures = [
0.00300,
0.00400,
0.00900,
0.01400,
0.02100,
0.02600,
0.03600,
0.05100,
0.09300,
0.15000,
0.20300,
0.30000,
0.40400,
0.50300,
0.60200,
0.70200,
0.80100,
0.90500,
1.00400,
]
A first way to do this is to replace the time coordinates by the pressure coordinate
(we first make a copy of the time coordinates for later use the original will be destroyed by the following operation)
Now we perform the replacement with this new coordinate:
c_pressures = scp.Coord(pressures, title="pressure", units="torr")
X.y = c_pressures
print(X.y)
Coord: [float64] torr (size: 19)
A second way is to affect several coordinates to the corresponding dimension. To do this, the simplest is to affect a list of coordinates instead of a single one:
X.y = [c_times, c_pressures]
print(X.y)
CoordSet: [_1:acquisition timestamp (GMT), _2:pressure]
By default, the current coordinate is the first one (here c_times ).
For example, it will be used for plotting:
prefs = X.preferences
prefs.figure.figsize = (7, 3)
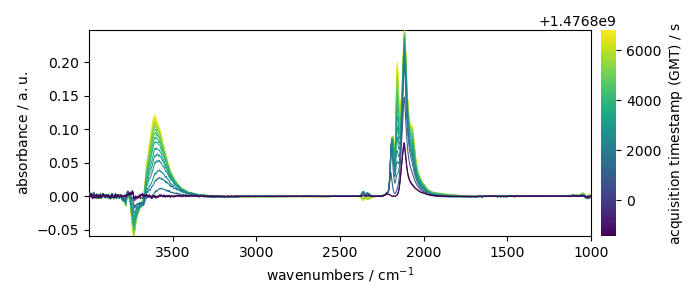
_ = X.plot(colorbar=True)
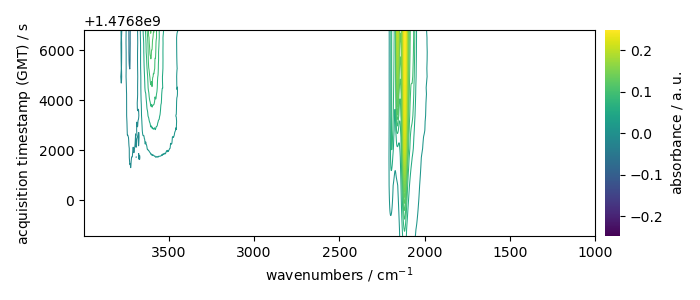
_ = X.plot_map(colorbar=True)
To seamlessly work with the second coordinates (pressures), we can change the default coordinate:
X.y.select(2) # to select coordinate `_2`
X.y.default
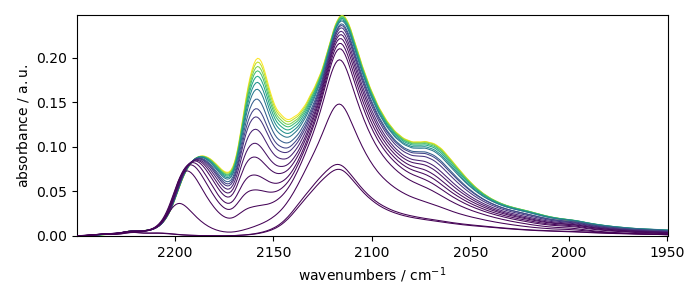
Let’s now plot the spectral range of interest. The default coordinate is now used:
X_ = X[:, 2250.0:1950.0]
print(X_.y.default)
_ = X_.plot()
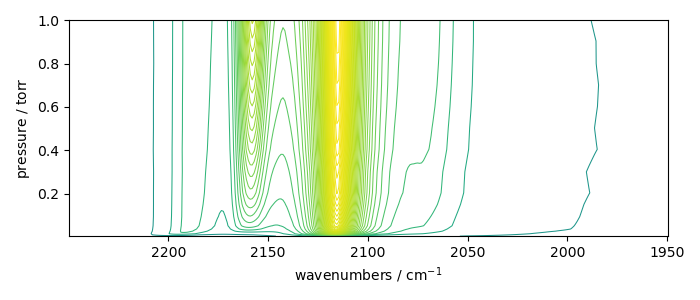
_ = X_.plot_map()
Coord: [float64] torr (size: 19)
The same can be done for the x coordinates.
Let’s take for instance row with index 10 of the previous dataset
row10 = X_[10].squeeze()
row10.plot()
print(row10.coordset)
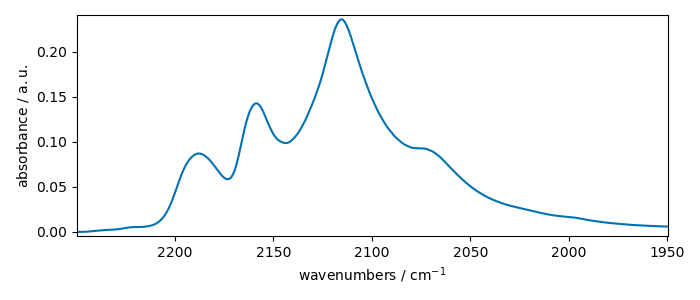
CoordSet: [x:wavenumbers]
Now we wants to add a coordinate with the wavelength instead of wavenumber.
c_wavenumber = row10.x.copy()
c_wavelength = row10.x.to("nanometer")
print(c_wavenumber, c_wavelength)
row10.x = [c_wavenumber, c_wavelength]
row10.x.select(2)
_ = row10.plot()
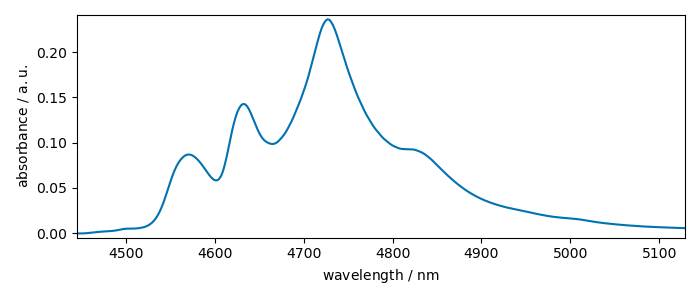
Coord: [float64] cm⁻¹ (size: 312) Coord: [float64] nm (size: 312)
This ends the example ! The following line can be uncommented if no plot shows when running the .py script
scp.show()
Total running time of the script: ( 0 minutes 1.565 seconds)