spectrochempy.PCA
- class PCA(*, log_level='WARNING', warm_start=False, iterated_power='auto', n_components=None, n_oversamples=10, power_iteration_normalizer='auto', random_state=None, scaled=False, standardized=False, svd_solver='auto', tol=0.0, whiten=False)[source]
Principal Component Anamysis (PCA).
The Principal Component Analysis analysis is using the
sklearn.decomposition.PCAmodel.- Parameters:
log_level (any of [
"INFO","DEBUG","WARNING","ERROR"], optional, default:"WARNING") – The log level at startup. It can be changed later on using theset_log_levelmethod or by changing thelog_levelattribute.warm_start (
bool, optional, default:False) – When fitting repeatedly on the same dataset, but for multiple parameter values (such as to find the value maximizing performance), it may be possible to reuse previous model learned from the previous parameter value, saving time.When
warm_startisTrue, the existing fitted model attributes is used to initialize the new model in a subsequent call tofit.iterated_power (an int or any of [‘auto’], optional, default:
'auto') – Number of iterations for the power method computed by svd_solver == ‘randomized’. Must be of range [0, infinity).n_components (any of [‘mle’] or an int or a float, optional, default:
None) – Number of components to keep. ifn_componentsis not set all components are kept:n_components == min(n_observations, n_features)
If
n_components == 'mle'andsvd_solver == 'full', Minka’s MLE is used to guess the dimension. Use ofn_components == 'mle'will interpretsvd_solver == 'auto'assvd_solver == 'full'. If0 < n_components < 1andsvd_solver == 'full', select the number of components such that the amount of variance that needs to be explained is greater than the percentage specified by n_components. Ifsvd_solver == 'arpack', the number of components must be strictly less than the minimum of n_features and n_observations. Hence, the None case results in:n_components == min(n_observations, n_features) - 1.
n_oversamples (
int, optional, default: 10) – This parameter is only relevant whensvd_solver="randomized". It corresponds to the additional number of random vectors to sample the range ofXso as to ensure proper conditioning. Seerandomized_svdfor more details.power_iteration_normalizer (any value of [
'auto','QR','LU','none'], optional, default:'auto') – Power iteration normalizer for randomized SVD solver. Not used by ARPACK. Seerandomized_svdfor more details.random_state (an int or a RandomState, optional, default:
None) – Used when the ‘arpack’ or ‘randomized’ solvers are used. Pass an int for reproducible results across multiple function calls.scaled (
bool, optional, default: False) – If True the data are scaled in the interval[0-1]: \(X' = (X - min(X)) / (max(X)-min(X))\).standardized (
bool, optional, default: False) – If True the data are scaled to unit standard deviation: \(X' = X / \sigma\).svd_solver (any value of [
'auto','full','arpack','randomized'], optional, default:'auto') – If auto : The solver is selected by a default policy based onX.shapeandn_components: if the input data is larger than 500x500 and the number of components to extract is lower than 80% of the smallest dimension of the data, then the more efficient ‘randomized’ method is enabled. Otherwise the exact full SVD is computed and optionally truncated afterwards. If full : run exact full SVD calling the standard LAPACK solver viascipy.linalg.svdand select the components by postprocessing If arpack : run SVD truncated to n_components calling ARPACK solver viascipy.sparse.linalg.svds. It requires strictly 0 < n_components < min(X.shape) If randomized : run randomized SVD by the method of Halko et al.tol (
float, optional, default: 0.0) – Tolerance for singular values computed by svd_solver == ‘arpack’. Must be of range [0.0, infinity).whiten (
bool, optional, default: False) – When True (False by default) thecomponents_vectors are multiplied by the square root of n_observations and then divided by the singular values to ensure uncorrelated outputs with unit component-wise variances. Whitening will remove some information from the transformed signal (the relative variance scales of the components) but can sometime improve the predictive accuracy of the downstream estimators by making their data respect some hard-wired assumptions.
See also
EFAPerform an Evolving Factor Analysis (forward and reverse).
FastICAPerform Independent Component Analysis with a fast algorithm.
IRISIntegral inversion solver for spectroscopic data.
MCRALSPerform MCR-ALS of a dataset knowing the initial \(C\) or \(S^T\) matrix.
NMFNon-Negative Matrix Factorization.
SIMPLISMASIMPLe to use Interactive Self-modeling Mixture Analysis.
SVDPerform a Singular Value Decomposition.
Initialize the BaseConfigurable class.
- Parameters:
log_level (int, optional) – The log level at startup. Default is logging.WARNING.
**kwargs (dict) – Additional keyword arguments for configuration.
Attributes Summary
Return the X input dataset (eventually modified by the model).
The
Yinput.NDDatasetwith components in feature space (n_components, n_features).traitlets.config.Configobject.Number of iterations for the power method computed by svd_solver == 'randomized'.
Return PCA loadings.
Return
logoutput.Number of components to keep. if
n_componentsis not set all components are kept::.This parameter is only relevant when
svd_solver="randomized".Object name
Power iteration normalizer for randomized SVD solver.
Used when the 'arpack' or 'randomized' solvers are used.
\(X' = (X - min(X)) / (max(X)-min(X))\).
Returns PCA scores.
\(X' = X / \sigma\).
If auto: The solver is selected by a default policy based on
X.shapeandn_components: if the input data is larger than 500x500 and the number of components to extract is lower than 80% of the smallest dimension of the data, then the more efficient 'randomized' method is enabled.Tolerance for singular values computed by svd_solver == 'arpack'.
When True (False by default) the
components_vectors are multiplied by the square root of n_observations and then divided by the singular values to ensure uncorrelated outputs with unit component-wise variances.Methods Summary
fit(X)Fit the PCA model on X.
fit_transform(X[, Y])Fit the model with
Xand apply the dimensionality reduction onX.get_components([n_components])Return the component's dataset: (selected n_components, n_features).
inverse_transform([X_transform])Transform data back to its original space.
parameters([replace, removed, default])Alias for
paramsmethod.params([default])Return current or default configuration values.
plot_merit([X, X_hat])Plot the input (
X), reconstructed (X_hat) and residuals.plot_score([scores, components])2D or 3D score plot of observations.
plot_scree([n_components])Scree plot of explained variance + cumulative variance by PCA.
plotmerit([X, X_hat])Plot the input (
X), reconstructed (X_hat) and residuals.printev([n_components])Print PCA figures of merit.
reconstruct([X_transform])Transform data back to its original space.
reduce([X])Apply dimensionality reduction to
X.reset()Reset configuration parameters to their default values.
scoreplot(self, *args, **kwargs)2D or 3D scoreplot of observations.
screeplot([n_components])Scree plot of explained variance + cumulative variance by PCA.
to_dict()Return config value in a dict form.
transform([X])Apply dimensionality reduction to
X.Attributes Documentation
- X
Return the X input dataset (eventually modified by the model).
- components
NDDatasetwith components in feature space (n_components, n_features).See also
get_componentsRetrieve only the specified number of components.
- config
traitlets.config.Configobject.
- iterated_power
Number of iterations for the power method computed by svd_solver == ‘randomized’. Must be of range [0, infinity).
- loadings
Return PCA loadings.
- log
Return
logoutput.
- n_components
Number of components to keep. if
n_componentsis not set all components are kept:n_components == min(n_observations, n_features)
If
n_components == 'mle'andsvd_solver == 'full', Minka’s MLE is used to guess the dimension. Use ofn_components == 'mle'will interpretsvd_solver == 'auto'assvd_solver == 'full'. If0 < n_components < 1andsvd_solver == 'full', select the number of components such that the amount of variance that needs to be explained is greater than the percentage specified by n_components. Ifsvd_solver == 'arpack', the number of components must be strictly less than the minimum of n_features and n_observations. Hence, the None case results in:n_components == min(n_observations, n_features) - 1.
- n_oversamples
This parameter is only relevant when
svd_solver="randomized". It corresponds to the additional number of random vectors to sample the range ofXso as to ensure proper conditioning. Seerandomized_svdfor more details.
- name
Object name
- power_iteration_normalizer
Power iteration normalizer for randomized SVD solver. Not used by ARPACK. See
randomized_svdfor more details.
- random_state
Used when the ‘arpack’ or ‘randomized’ solvers are used. Pass an int for reproducible results across multiple function calls.
- scaled
\(X' = (X - min(X)) / (max(X)-min(X))\).
- Type:
If True the data are scaled in the interval
[0-1]
- scores
Returns PCA scores.
- standardized
\(X' = X / \sigma\).
- Type:
If True the data are scaled to unit standard deviation
- svd_solver
If auto: The solver is selected by a default policy based on
X.shapeandn_components: if the input data is larger than 500x500 and the number of components to extract is lower than 80% of the smallest dimension of the data, then the more efficient ‘randomized’ method is enabled. Otherwise the exact full SVD is computed and optionally truncated afterwards. If full : run exact full SVD calling the standard LAPACK solver viascipy.linalg.svdand select the components by postprocessing If arpack : run SVD truncated to n_components calling ARPACK solver viascipy.sparse.linalg.svds. It requires strictly 0 < n_components < min(X.shape) If randomized : run randomized SVD by the method of Halko et al.
- tol
Tolerance for singular values computed by svd_solver == ‘arpack’. Must be of range [0.0, infinity).
- whiten
When True (False by default) the
components_vectors are multiplied by the square root of n_observations and then divided by the singular values to ensure uncorrelated outputs with unit component-wise variances. Whitening will remove some information from the transformed signal (the relative variance scales of the components) but can sometime improve the predictive accuracy of the downstream estimators by making their data respect some hard-wired assumptions.
Methods Documentation
- fit(X)[source]
Fit the PCA model on X.
- Parameters:
X (
NDDatasetor array-like of shape (n_observations, n_features)) – Training data.- Returns:
self – The fitted instance itself.
See also
fit_transformFit the model with an input dataset
Xand apply the dimensionality reduction onX.fit_reduceAlias of
fit_transform(Deprecated).
- fit_transform(X, Y=None, **kwargs)[source]
Fit the model with
Xand apply the dimensionality reduction onX.- Parameters:
X (
NDDatasetor array-like of shape (n_observations, n_features)) – Training data.Y (any) – Depends on the model.
**kwargs (keyword parameters, optional) – See Other Parameters.
- Returns:
NDDataset– Dataset with shape (n_observations, n_components).- Other Parameters:
n_components (
int, optional) – The number of components to use for the reduction. If not given the number of components is eventually the one specified or determined in thefitprocess.
- get_components(n_components=None)
Return the component’s dataset: (selected n_components, n_features).
- Parameters:
n_components (
int, optional, default:None) – The number of components to keep in the output dataset. IfNone, all calculated components are returned.- Returns:
NDDataset– Dataset with shape (n_components, n_features)
- inverse_transform(X_transform=None, **kwargs)
Transform data back to its original space.
In other words, return an input
X_originalwhose reduce/transform would beX_transform.- Parameters:
X_transform (array-like of shape (n_observations, n_components), optional) – Reduced
Xdata, wheren_observationsis the number of observations andn_componentsis the number of components. IfX_transformis not provided, a transform ofXprovided infitis performed first.**kwargs (keyword parameters, optional) – See Other Parameters.
- Returns:
NDDataset– Dataset with shape (n_observations, n_features).- Other Parameters:
n_components (
int, optional) – The number of components to use for the reduction. If not given the number of components is eventually the one specified or determined in thefitprocess.
See also
reconstructAlias of inverse_transform (Deprecated).
- parameters(replace="params", removed="0.8.0") def parameters(self, default=False)[source]
Alias for
paramsmethod.Deprecated since version 0.8.0: Use
paramsinstead.
- plot_merit(X=None, X_hat=None, **kwargs)[source]
Plot the input (
X), reconstructed (X_hat) and residuals.\(X\) and \(\hat{X}\) can be passed as arguments. If not, the
Xattribute is used for \(X`and :math:\)hat{X}`is computed by theinverse_transformmethod- Parameters:
X (
NDDataset, optional) – Original dataset. If is not provided (default), theXattribute is used and X_hat is computed usinginverse_transform.X_hat (
NDDataset, optional) – Inverse transformed dataset. ifXis provided,X_hatmust also be provided as compuyed externally.
- Returns:
Axes– Matplotlib subplot axe.- Other Parameters:
exp_c (color, colormap, or list of colors, optional) – Color(s) for experimental spectra. - None: use unified semantic resolver (auto-detect categorical/sequential) - Single color: use for all experimental spectra - Colormap name/object: sample colors from colormap - List/tuple: use as explicit color cycle
calc_c (color, colormap, or list of colors, optional) – Color(s) for calculated spectra. - None: use default blue “#2a6fbb” - Single color: use for all calculated spectra - Colormap name/object: sample colors from colormap - List/tuple: use as explicit color cycle
resid_c (color, colormap, or list of colors, optional) – Color(s) for residual spectra. - None: use default grey “0.4” - Single color: use for all residual spectra - Colormap name/object: sample colors from colormap - List/tuple: use as explicit color cycle
exp_linestyle (str, optional) – Line style for experimental spectra. Default: “-“.
calc_linestyle (str, optional) – Line style for calculated spectra. Default: “–“.
resid_linestyle (str, optional) – Line style for residual spectra. Default: “-“.
exp_linewidth (float, optional) – Line width for experimental spectra. Default: 1.2.
calc_linewidth (float, optional) – Line width for calculated spectra. Default: 1.0.
resid_linewidth (float, optional) – Line width for residual spectra. Default: 1.0.
min_contrast (float, optional) – Minimum contrast ratio for sequential colormaps. Default: 1.5.
offset (
float, optional, default:None) – Specify the separation (in percent) between the \(X\) , \(X_hat\) and \(E\).nb_traces (
intor'all', optional) – Number of lines to display. Default is'all'.**others (Other keywords parameters) – Parameters passed to the internal
plotmethod of theXdataset.
- plot_score(scores=None, components=(1, 2), **kwargs)[source]
2D or 3D score plot of observations.
Plots the projection of each observation/spectrum onto the span of two or three selected principal components.
- Parameters:
scores (NDDataset or tuple, optional) – Scores dataset to plot. If None, uses
self.scores. Pass a modified scores dataset (e.g., with custom labels) to use those labels in the plot.Note: If a tuple or list is passed as the first positional argument, it is interpreted as
componentsfor backward compatibility.components (tuple of int, optional) – Principal components to plot (1-based indexing). Length 2 for 2D plot, length 3 for 3D plot. Default: (1, 2).
**kwargs – Additional keyword arguments passed to
plot_score. Seeplot_scorefor available options:cmap: Colormap for coloring points.color: Fixed color or color values for each point.color_mapping: “index” (default) or “labels” - how to map colors.show_labels: If True, annotate points with labels.labels_column: Column index in scores.y.labels (0-based).ax: Axes to plot on.show: Whether to display the figure.
- Returns:
matplotlib.axes.Axes – The matplotlib axes.
See also
plot_scoreStandalone score plot function.
Examples
>>> import spectrochempy as scp >>> X = scp.read("irdata/nh4y-activation.spg") >>> pca = scp.PCA(n_components=5) >>> pca.fit(X) >>> ax = pca.plot_score((1, 2), show=False)
With custom labels:
>>> scores = pca.transform() >>> scores.y.labels = custom_labels >>> ax = pca.plot_score(scores=scores, show_labels=True)
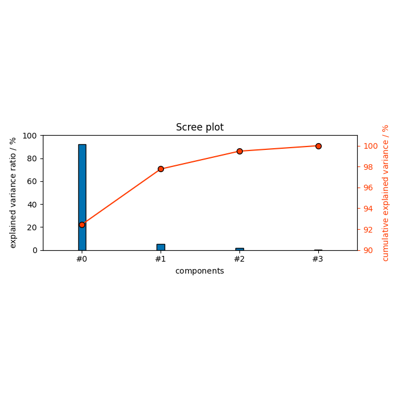
- plot_scree(n_components=None, **kwargs)[source]
Scree plot of explained variance + cumulative variance by PCA.
Explained variance by each PC is plotted as a bar graph (left y axis) and cumulative explained variance is plotted as a line with markers (right y axis).
- Parameters:
n_components (int, optional) – Number of components to plot. If None, plots all components.
**kwargs – Additional keyword arguments passed to
plot_scree. Seeplot_screefor available options (e.g.,bar_color,line_color,title,ax,show).
- Returns:
matplotlib.axes.Axes – The primary axes (left y-axis with bars).
See also
plot_screeStandalone scree plot function.
Examples
>>> import spectrochempy as scp >>> X = scp.read("irdata/nh4y-activation.spg") >>> pca = scp.PCA(n_components=5) >>> pca.fit(X) >>> ax = pca.plot_scree(show=False)
- plotmerit(X=None, X_hat=None, **kwargs)[source]
Plot the input (
X), reconstructed (X_hat) and residuals.\(X\) and \(\hat{X}\) can be passed as arguments. If not, the
Xattribute is used for \(X`and :math:\)hat{X}`is computed by theinverse_transformmethod- Parameters:
X (
NDDataset, optional) – Original dataset. If is not provided (default), theXattribute is used and X_hat is computed usinginverse_transform.X_hat (
NDDataset, optional) – Inverse transformed dataset. ifXis provided,X_hatmust also be provided as compuyed externally.
- Returns:
Axes– Matplotlib subplot axe.- Other Parameters:
exp_c (color, colormap, or list of colors, optional) – Color(s) for experimental spectra. - None: use unified semantic resolver (auto-detect categorical/sequential) - Single color: use for all experimental spectra - Colormap name/object: sample colors from colormap - List/tuple: use as explicit color cycle
calc_c (color, colormap, or list of colors, optional) – Color(s) for calculated spectra. - None: use default blue “#2a6fbb” - Single color: use for all calculated spectra - Colormap name/object: sample colors from colormap - List/tuple: use as explicit color cycle
resid_c (color, colormap, or list of colors, optional) – Color(s) for residual spectra. - None: use default grey “0.4” - Single color: use for all residual spectra - Colormap name/object: sample colors from colormap - List/tuple: use as explicit color cycle
exp_linestyle (str, optional) – Line style for experimental spectra. Default: “-“.
calc_linestyle (str, optional) – Line style for calculated spectra. Default: “–“.
resid_linestyle (str, optional) – Line style for residual spectra. Default: “-“.
exp_linewidth (float, optional) – Line width for experimental spectra. Default: 1.2.
calc_linewidth (float, optional) – Line width for calculated spectra. Default: 1.0.
resid_linewidth (float, optional) – Line width for residual spectra. Default: 1.0.
min_contrast (float, optional) – Minimum contrast ratio for sequential colormaps. Default: 1.5.
offset (
float, optional, default:None) – Specify the separation (in percent) between the \(X\) , \(X_hat\) and \(E\).nb_traces (
intor'all', optional) – Number of lines to display. Default is'all'.**others (Other keywords parameters) – Parameters passed to the internal
plotmethod of theXdataset.
- printev(n_components=None)[source]
Print PCA figures of merit.
Prints eigenvalues and explained variance for all or first n_pc PC’s.
- Parameters:
n_components (int, optional) – The number of components to print.
- reconstruct(X_transform=None, **kwargs)[source]
Transform data back to its original space.
In other words, return an input
X_originalwhose reduce/transform would beX_transform.- Parameters:
X_transform (array-like of shape (n_observations, n_components), optional) – Reduced
Xdata, wheren_observationsis the number of observations andn_componentsis the number of components. IfX_transformis not provided, a transform ofXprovided infitis performed first.**kwargs (keyword parameters, optional) – See Other Parameters.
- Returns:
NDDataset– Dataset with shape (n_observations, n_features).- Other Parameters:
n_components (
int, optional) – The number of components to use for the reduction. If not given the number of components is eventually the one specified or determined in thefitprocess.
See also
reconstructAlias of inverse_transform (Deprecated).
Notes
Deprecated in version 0.6.
- reduce(X=None, **kwargs)[source]
Apply dimensionality reduction to
X.- Parameters:
X (
NDDatasetor array-like of shape (n_observations, n_features), optional) – New data, where n_observations is the number of observations and n_features is the number of features. if not provided, the input dataset of thefitmethod will be used.**kwargs (keyword parameters, optional) – See Other Parameters.
- Returns:
NDDataset– Dataset with shape (n_observations, n_components).- Other Parameters:
n_components (
int, optional) – The number of components to use for the reduction. If not given the number of components is eventually the one specified or determined in thefitprocess.
Notes
Deprecated in version 0.6.
- scoreplot(self, *args, **kwargs)[source]
2D or 3D scoreplot of observations.
Deprecated since version 0.7.4: Use
plot_scoreinstead.- Parameters:
*args – Positional arguments. Accepts: - (i, j) or (i, j, k): component indices (1-based) - (scores, i, j): NDDataset and component indices
**kwargs – Additional keyword arguments passed to
plot_score.
- Returns:
matplotlib.axes.Axes – The matplotlib axes.
- screeplot(n_components=None, **kwargs)[source]
Scree plot of explained variance + cumulative variance by PCA.
Deprecated since version 0.7.4: Use
plot_screeinstead.- Parameters:
n_components (int, optional) – Number of components to plot.
**kwargs – Additional keyword arguments (ignored, for backward compatibility).
- Returns:
matplotlib.axes.Axes – The primary axes.
- transform(X=None, **kwargs)
Apply dimensionality reduction to
X.- Parameters:
X (
NDDatasetor array-like of shape (n_observations, n_features), optional) – New data, where n_observations is the number of observations and n_features is the number of features. if not provided, the input dataset of thefitmethod will be used.**kwargs (keyword parameters, optional) – See Other Parameters.
- Returns:
NDDataset– Dataset with shape (n_observations, n_components).- Other Parameters:
n_components (
int, optional) – The number of components to use for the reduction. If not given the number of components is eventually the one specified or determined in thefitprocess.
Examples using spectrochempy.PCA