Reading datasets
In this example, we show the use of the generic read method to create dataset
either from local or remote files.
First we need to import the spectrochempy API package
import spectrochempy as scp
Import dataset from local files
Read a IR data recorded in Omnic format (.spg extension).
We just pass the file name as parameter.
NDDataset: [float64] a.u. (shape: (y:55, x:5549))[nh4y-activation]
Summary
author
:
runner@runnervm3jyl0
created
:
2026-05-31 15:53:35+00:00
description
:
Omnic title: NH4Y-activation.SPG
Omnic filename: /home/runner/.spectrochempy/testdata/irdata/nh4y-activation.spg
history
:
2026-05-31 15:53:35+00:00> Imported from spg file /home/runner/.spectrochempy/testdata/irdata/nh4y-activation.spg.
2026-05-31 15:53:35+00:00> Sorted by date
Data
[[ 2.057 2.061 ... 2.013 2.012]
[ 2.033 2.037 ... 1.913 1.911]
...
[ 1.794 1.791 ... 1.198 1.198]
[ 1.816 1.815 ... 1.24 1.238]] a.u.
Dimension `x`
coordinates
:
[ 6000 5999 ... 650.9 649.9] cm⁻¹
Dimension `y`
title
:
acquisition timestamp (GMT)
coordinates
:
[1.468e+09 1.468e+09 ... 1.468e+09 1.468e+09] s
[[ 2016-07-06 19:03:14+00:00 2016-07-06 19:13:14+00:00 ... 2016-07-07 04:03:17+00:00 2016-07-07 04:13:17+00:00]
[ vz0466.spa, Wed Jul 06 21:00:38 2016 (GMT+02:00) vz0467.spa, Wed Jul 06 21:10:38 2016 (GMT+02:00) ...
vz0520.spa, Thu Jul 07 06:00:41 2016 (GMT+02:00) vz0521.spa, Thu Jul 07 06:10:41 2016 (GMT+02:00)]]
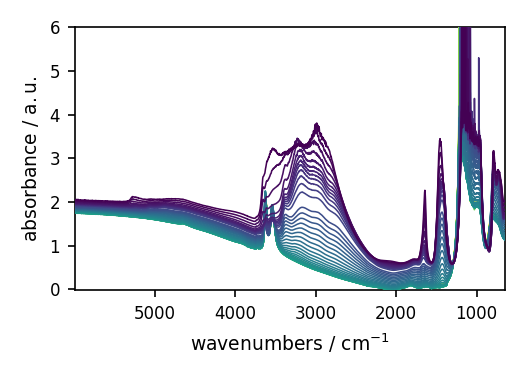
When using read, we can pass filename as a str or a Path object.
Note that is the file is not found in the current working directory, SpectroChemPy
will try to find it in the datadir directory defined in preferences :
PosixPath('/home/runner/.spectrochempy/testdata')
If the supplied argument is a directory, then the whole directory is read at once.
By default, the different files will be merged along the first dimension (y).
However, for this to work, the second dimension (x) must be compatible (same size)
or else a WARNING appears. To avoid the warning and get individual spectra, you can
set merge to False .
dataset_list = scp.read("irdata", merge=False)
dataset_list
List (len=3, type=NDDataset)
0: NDDataset: [float64] a.u. (shape: (y:19, x:3112))[CO@Mo_Al2O3]
Summary
author
:
runner@runnervm3jyl0
created
:
2026-05-31 15:53:36+00:00
description
:
Omnic title: Group sust Mo_Al2O3_base line.SPG
Omnic filename: /home/runner/.spectrochempy/testdata/irdata/CO@Mo_Al2O3.SPG
history
:
2026-05-31 15:53:36+00:00> Imported from spg file /home/runner/.spectrochempy/testdata/irdata/CO@Mo_Al2O3.SPG.
2026-05-31 15:53:36+00:00> Sorted by date
Data
[[0.0008032 3.788e-05 ... 0.0003027 0.0003745]
[-3.608e-05 -0.0001981 ... 0.0003089 0.00117]
...
[0.0008357 -0.0001387 ... -0.0005221 -0.001121]
[0.0005655 -0.000116 ... -0.00057 -0.0006307]] a.u.
Dimension `x`
coordinates
:
[ 4000 3999 ... 1001 999.9] cm⁻¹
Dimension `y`
title
:
acquisition timestamp (GMT)
coordinates
:
[1.477e+09 1.477e+09 ... 1.477e+09 1.477e+09] s
[[ 2016-10-18 13:49:35+00:00 2016-10-18 13:54:06+00:00 ... 2016-10-18 16:01:33+00:00 2016-10-18 16:06:37+00:00]
[ *Résultat de Soustraction:04_Mo_Al2O3_calc_0.003torr_LT_after sulf_Oct 18 15:46:42 2016 (GMT+02:00)
*Résultat de Soustraction:04_Mo_Al2O3_calc_0.004torr_LT_after sulf_Oct 18 15:51:12 2016 (GMT+02:00) ...
*Résultat de Soustraction:04_Mo_Al2O3_calc_0.905torr_LT_after sulf_Oct 18 17:58:42 2016 (GMT+02:00)
*Résultat de Soustraction:04_Mo_Al2O3_calc_1.004torr_LT_after sulf_Oct 18 18:03:41 2016 (GMT+02:00)]]
1: NDDataset: [float64] a.u. (shape: (y:55, x:5549))[nh4y-activation]
Summary
author
:
runner@runnervm3jyl0
created
:
2026-05-31 15:53:36+00:00
description
:
Omnic title: NH4Y-activation.SPG
Omnic filename: /home/runner/.spectrochempy/testdata/irdata/nh4y-activation.spg
history
:
2026-05-31 15:53:36+00:00> Imported from spg file /home/runner/.spectrochempy/testdata/irdata/nh4y-activation.spg.
2026-05-31 15:53:36+00:00> Sorted by date
Data
[[ 2.057 2.061 ... 2.013 2.012]
[ 2.033 2.037 ... 1.913 1.911]
...
[ 1.794 1.791 ... 1.198 1.198]
[ 1.816 1.815 ... 1.24 1.238]] a.u.
Dimension `x`
coordinates
:
[ 6000 5999 ... 650.9 649.9] cm⁻¹
Dimension `y`
title
:
acquisition timestamp (GMT)
coordinates
:
[1.468e+09 1.468e+09 ... 1.468e+09 1.468e+09] s
[[ 2016-07-06 19:03:14+00:00 2016-07-06 19:13:14+00:00 ... 2016-07-07 04:03:17+00:00 2016-07-07 04:13:17+00:00]
[ vz0466.spa, Wed Jul 06 21:00:38 2016 (GMT+02:00) vz0467.spa, Wed Jul 06 21:10:38 2016 (GMT+02:00) ...
vz0520.spa, Thu Jul 07 06:00:41 2016 (GMT+02:00) vz0521.spa, Thu Jul 07 06:10:41 2016 (GMT+02:00)]]
2: NDDataset: [float64] unitless (shape: (y:1, x:3736))[IR]
Summary
author
:
runner@runnervm3jyl0
created
:
2026-05-31 15:53:36+00:00
description
:
"name" read from .csv file
history
:
2026-05-31 15:53:36+00:00> Read from .csv file
Data
[[-0.09079 3.547 ... 4.317 -0.09079]]
Dimension `x`
coordinates
:
[ 399.2 400.2 ... 4000 4001]
to get full details on the parameters that can be used, look at the API documentation:
spectrochempy.read .
Import dataset from remote files
To download and read file from remote server you can use urls.
dataset_list = scp.read("http://www.eigenvector.com/data/Corn/corn.mat")
In this case the matlab data contains 7 arrays that have been automatically
transformed to NDDataset .
mp6spec : (240, 700)
m5nbs : (3, 700)
mp6nbs : (8, 700)
propvals : (80, 4)
The eigenvector.com website contains the same data in a
compressed (zipped) format:
corn.mat_.zip .
This can also be used by the read method.
dataset_list = scp.read(
"https://eigenvector.com/wp-content/uploads/2019/06/corn.mat_.zip"
)
dataset_list
List (len=4, type=NDDataset)
0: NDDataset: [float64] unitless (shape: (y:240, x:700))[mp6spec]
Summary
created
:
2026-05-31 15:53:37+00:00
description
:
Concatenation of 3 datasets:
( m5spec, mp5spec, mp6spec )
history
:
2026-05-31 15:53:37+00:00> Created by concatenate
2026-05-31 15:53:37+00:00> Merged from several files
Data
[[ 0.04449 0.04438 ... 0.7309 0.7306]
[-0.01244 -0.01251 ... 0.6849 0.6844]
...
[-0.003701 -0.003818 ... 0.7128 0.7121]
[-0.01526 -0.01538 ... 0.7028 0.7021]]
Dimension `x`
coordinates
:
[ 1100 1102 ... 2496 2498]
1: NDDataset: [float64] unitless (shape: (y:3, x:700))[m5nbs]
Summary
created
:
2026-05-31 15:53:37+00:00
description
:
Concatenation of 1 datasets:
( m5nbs )
history
:
2026-05-31 15:53:37+00:00> Created by concatenate
2026-05-31 15:53:37+00:00> Merged from several files
Data
[[ 0.134 0.132 ... 0.09665 0.09771]
[ 0.1374 0.1354 ... 0.09898 0.09999]
[ 0.1437 0.1416 ... 0.1037 0.1048]]
Dimension `x`
coordinates
:
[ 1100 1102 ... 2496 2498]
2: NDDataset: [float64] unitless (shape: (y:8, x:700))[mp6nbs]
Summary
created
:
2026-05-31 15:53:37+00:00
description
:
Concatenation of 2 datasets:
( mp5nbs, mp6nbs )
history
:
2026-05-31 15:53:37+00:00> Created by concatenate
2026-05-31 15:53:37+00:00> Merged from several files
Data
[[ 0.06679 0.0645 ... 0.03636 0.03724]
[ 0.05404 0.0524 ... 0.02147 0.02236]
...
[ 0.06115 0.05901 ... 0.03013 0.03099]
[ 0.05311 0.05144 ... 0.02195 0.02284]]
Dimension `x`
coordinates
:
[ 1100 1102 ... 2496 2498]
3: NDDataset: [float64] unitless (shape: (y:80, x:4))[propvals]
Summary
created
:
2026-05-31 15:53:37+00:00
description
:
Concatenation of 1 datasets:
( propvals )
history
:
2026-05-31 15:53:37+00:00> Created by concatenate
2026-05-31 15:53:37+00:00> Merged from several files
Data
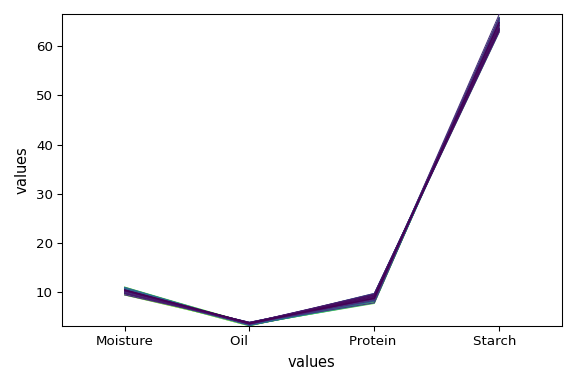
[[ 10.45 3.687 8.746 64.84]
[ 10.41 3.72 8.658 64.85]
...
[ 10.59 3.176 8.132 65.21]
[ 10.98 3.328 8.428 64.85]]
Dimension `x`
labels
:
[ Moisture Oil Protein Starch ]
Plot each of the datasets
dataset_list[-1].plot()
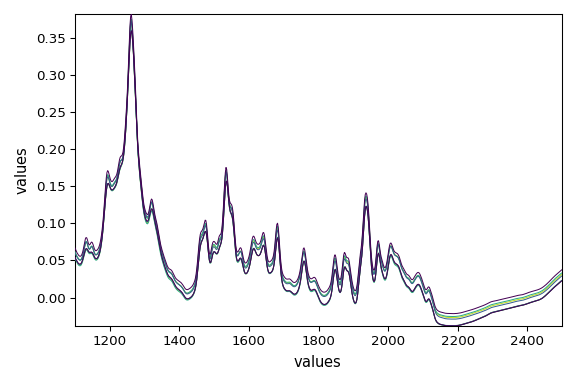
dataset_list[-2].plot()
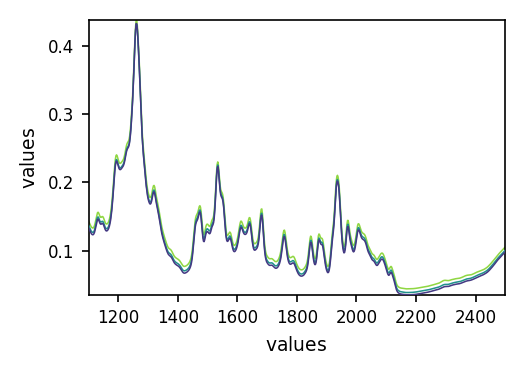
dataset_list[-3].plot()
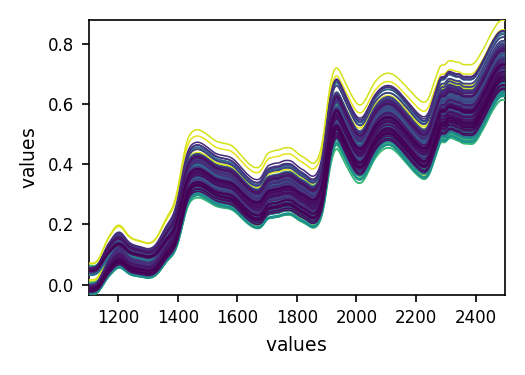
dataset_list[-4].plot()
<Axes: xlabel='values $\\mathrm{}$', ylabel='values $\\mathrm{}$'>
This ends the example ! The following line can be uncommented if no plot shows when
running the .py script with python
Total running time of the script: (0 minutes 2.495 seconds)