Note
Go to the end to download the full example code.
FastICA example
Import the spectrochempy API package
import spectrochempy as scp
import numpy as np
Independent component analysis (ICA) is a computational method for separating a multivariate signal such as spectra into additive components. This is done by assuming that at most one subcomponent is Gaussian and that the components are statistically independents from each other.
# Load, prepare and plot the dataset
# ----------------------------------
# Here we use a dataset from :cite:t:`jaumot:2005`
X = scp.read("matlabdata/als2004dataset.MAT")[-1]
X.title = "absorbance"
X.units = "absorbance"
X.set_coordset(
np.arange(X.shape[0], dtype="float"), None
) # y coordinates as floats to trigger sequential colormap
X.y.title = "elution time"
X.y.units = "min"
X.x.title = "wavelength"
X.plot()
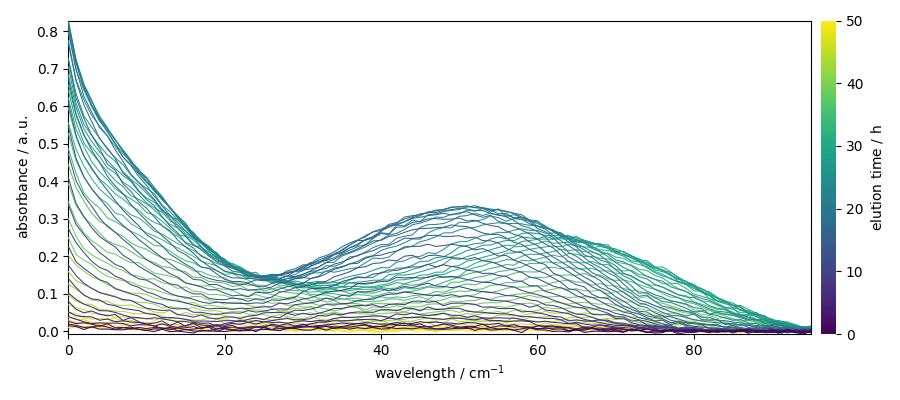
<Axes: xlabel='wavelength $\\mathrm{}$', ylabel='absorbance $\\mathrm{/\\ \\mathrm{a.u.}}$'>
Create and fit a FastICA object
As argument of the object constructor we define log_level to "INFO" to
obtain verbose output during fit, and we set the number of component to use at 4.
ica = scp.FastICA(n_components=4, log_level="INFO")
ica.fit(X)
<spectrochempy.analysis.decomposition.fast_ica.FastICA object at 0x7fa52b40eba0>
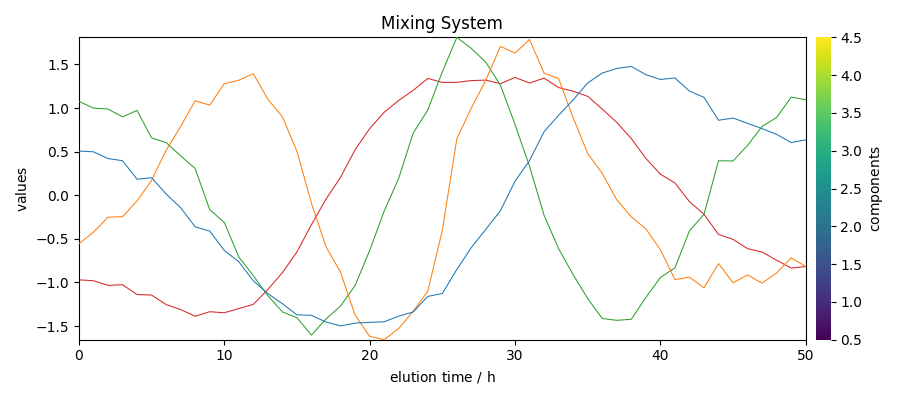
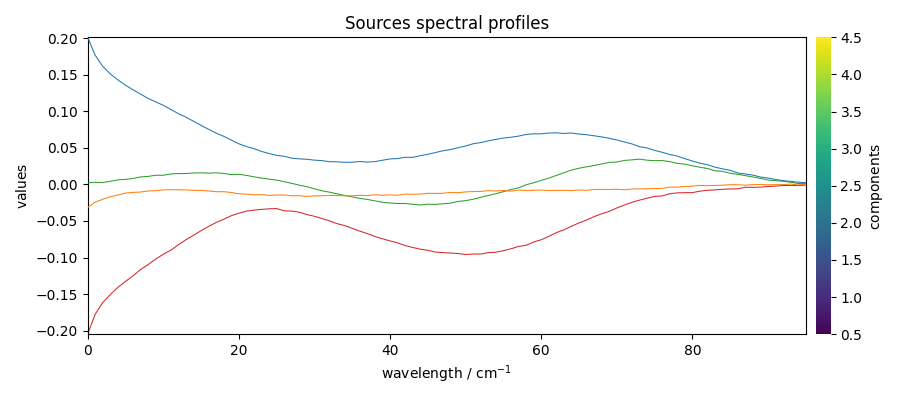
Get the mixing system and source spectral profiles
The mixing system \(A\) and the source spectral profiles \(S^T\) can be obtained as follows (the Sklearn equivalents - also valid with Scpy - are indicated as comments
Plot them
Reconstruct the dataset
The dataset can be reconstructed from these matrices and the mean:
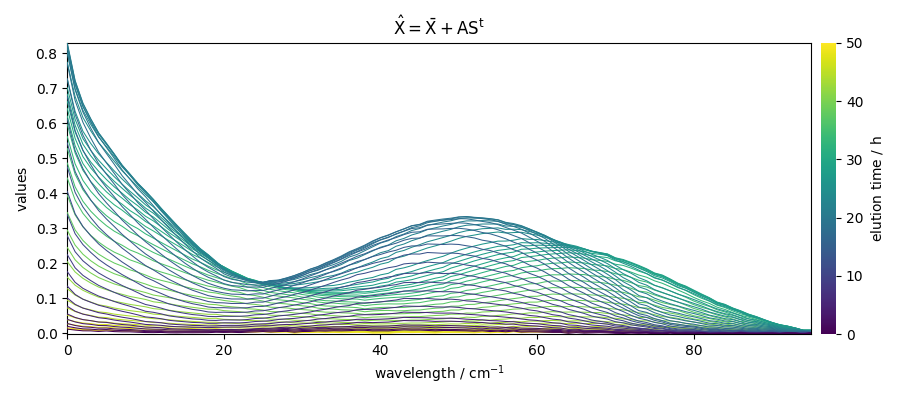
Or using the transform() method:
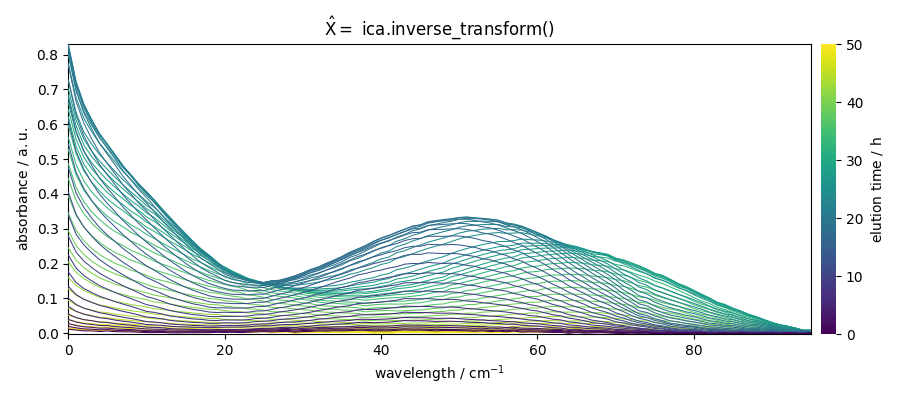
X_hat_b = ica.inverse_transform()
_ = X_hat_b.plot(title=r"$\hat{X} =$ ica.inverse_transform()")
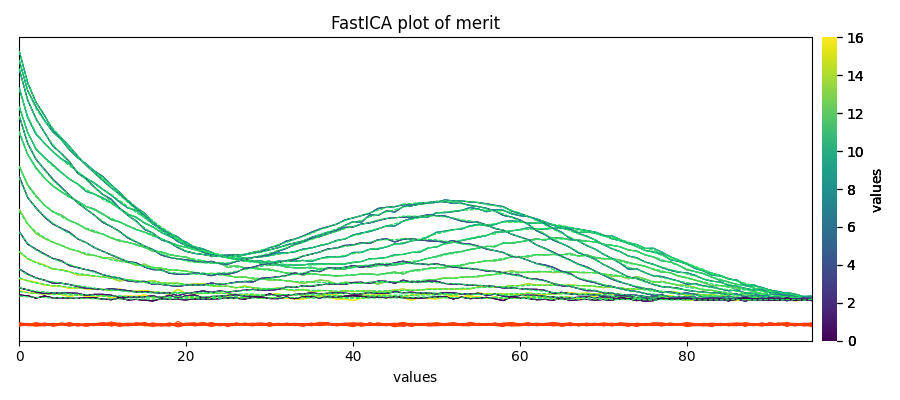
Finally, the quality of the reconstriction can be checked by plotmerit()
_ = ica.plotmerit(nb_traces=15)
This ends the example ! The following line can be uncommented if no plot shows when running the .py script with python
# scp.show()
Total running time of the script: (0 minutes 0.884 seconds)